|
10/11/2023 0 Comments 10x chromium read lengthWhich seems to be comparable to a CellRanger output summary for a sample that was mapped earlier. % of reads mapped to too many loci | 0.04% Number of reads mapped to too many loci | 30684 % of reads mapped to multiple loci | 4.33% Number of reads mapped to multiple loci | 3534767 Number of splices: Non-canonical | 280490 Number of splices: Annotated (sjdb) | 29052571 The final log out are as follows and indicate a good percentage of uniquely mapped reads. Thanks! I tried with putting the -soloBarcodeReadLength option in STAR to be 150 and there was no problem then. In CellRanger the length of reads is R1 is not problematic as there are options to specify trimming or the tool just ignores the rest of the reads. 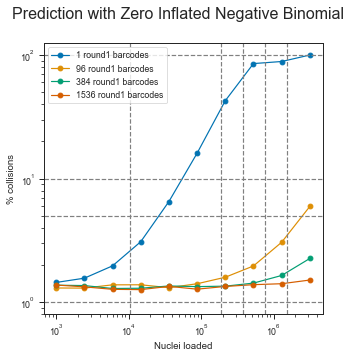
Do I need to trim the R1 reads to a length of 28bp before alignment? or (b) Should I just specify the -soloBarcodeReadLength option in STAR to be 150? If UMI+CB length is not equal to the barcode read length, specify barcode read length with -soloBarcodeReadLength" "SOLUTION: make sure that the barcode read is the last file in -readFilesIn, and check that it has the correct formatting Sequence=GAGGCAAGTGGCAGATCGTTTCAACATTGTTCCTGCGCAACACAGAATAGAGAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAA"Īs per STAR, the solution hint provided is the following: 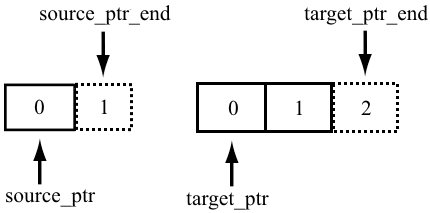
I have checked the FASTQ file for Read 1 and see that it is full 150bp. However, I am getting the following error and hope that some of you can help: EXITING because of FATAL ERROR in input read file: the total length of barcode sequence is 150 not equal to expected 28 I have some 10x v3 single cell rna seq fastq files that I am trying to map to human genome using STAR aligner.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
